class: center, middle, inverse, title-slide .title[ # Making your own R package … ] .subtitle[ ## will transform your life ] .author[ ### Elie Gurarie ] .date[ ### 2026-04-13 ] --- class: inverse, center, middle <style type="text/css"> .remark-slide-content { font-size: 20px; } </style> # Why build an R package? --- .pull-left-40[ ## Why!? - Conquer and permanently tame confusing folder soups of data and R scripts - Make code: - **ultra-compact** - **well-documented** - **highly replicable** - Dramatically shorten time to get back on track ultimately: - (truly) publish tools and methods ] .pull-right-60[ <br><br> <img src='images/version_control.gif' width='100%'/> ] --- ## Sonoran Pronghorn .pull-left-60[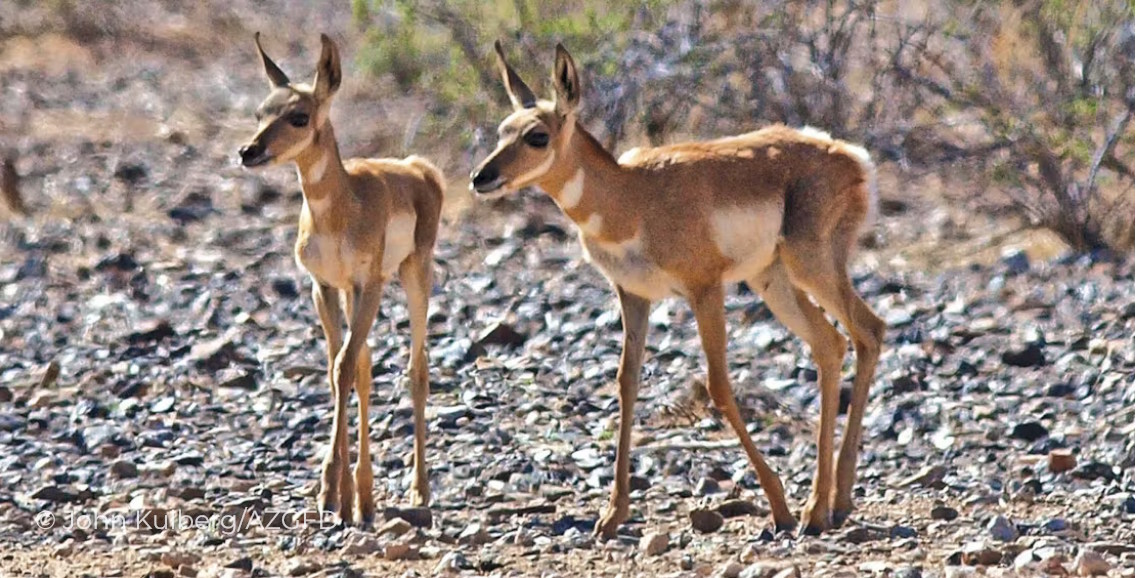  ] .pull-right-40[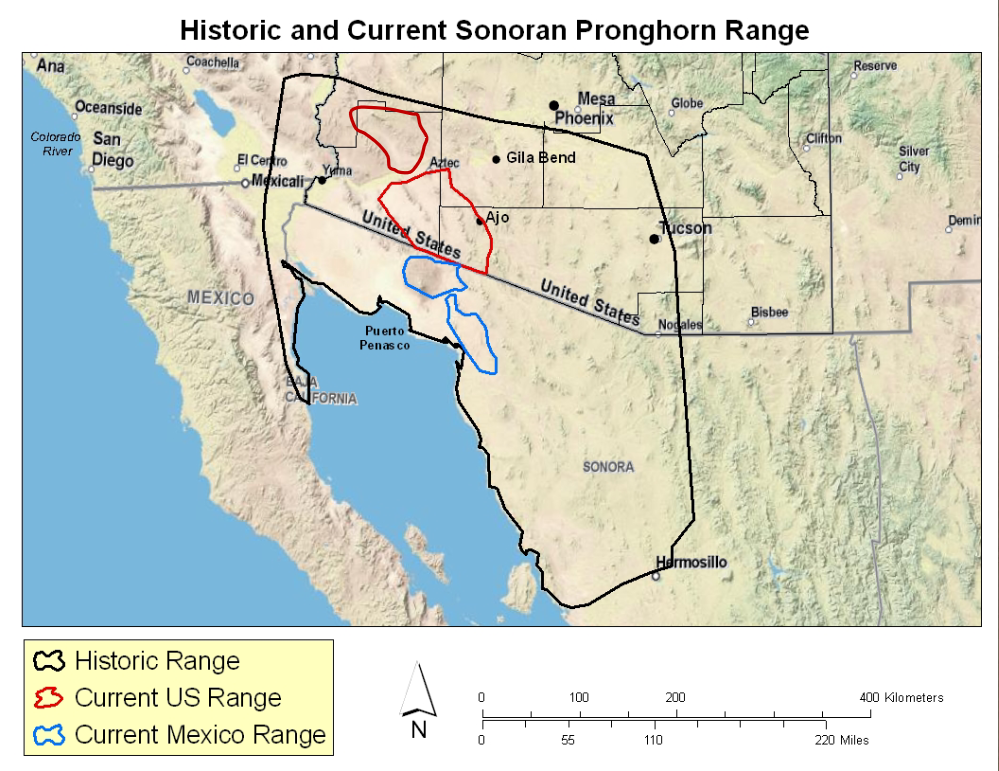  ] --- ## Processing before: (this is a small snippet of a data processing nightmare) .pull-left.small[ ``` r gps.dir <- "data/SonoranPronghorn/Locations_GPSCollarTelemetry/" pronghorn <- read.csv(paste0(gps.dir, f.v1[i])) %>% processRaw_v1(id = id.v1[i], filename = f.v1[i]) pronghorn.sf <- st_as_sf(df.raw, coords = c("ECEF_X..m.", "ECEF_Y..m.", "ECEF_Z..m.")) %>% st_set_crs(4978) %>% st_transform(4326) %>% st_coordinates with(df.raw, data.frame( File = filename, ID = CollarID, DateTime = mdy_hms(paste(UTC_Date, UTC_Time)), Latitude = ll[,"Y"], Longitude = ll[,"X"], Elevation = ll[,"Z"])) %>% subset(!is.na(DateTime)) ``` ] .pull-right.small[ ``` r # important step, need to convert from windows-1252 to UTF8 in order to read: # find *.csv -exec sh -c "iconv -f Windows-1252 -t UTF8 {} > {}v2" \; f.v2 <- f[grepl("GPS_Collar", f)] pronghorn_gps_v2 <- data.frame() for(i in 1:length(f.v2)){ if(f.v2[i] != badf){ print(f.v2[i]) df <- read.csv(paste0(gps.dir,"encoded/",f.v2[i])) %>% subset(!is.na(ECEF_X..m.)) %>% processRaw_v2(filename = f.v2[i]) pronghorn_gps_v2 <- rbind(pronghorn_gps_v2, df) } } ``` ] --- ## Processing now ``` r require(pronghorn) data("pronghorn_gps") head(pronghorn_gps) ``` ``` footnotesize ## File ID ## 17101 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## 18101 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## 19101 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## 20101 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## 21105 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## 22103 GPS_Collar_28269_Animal_NA_LastDataPullDate_20180822.csv 28269 ## DateTime Latitude Longitude Elevation ## 17101 2017-12-07 02:00:12 32.37972 -112.9905 457.5550 ## 18101 2017-12-07 13:00:38 32.37970 -112.9904 451.3434 ## 19101 2017-12-08 02:00:25 32.03507 -112.8549 583.3112 ## 20101 2017-12-08 13:00:39 32.03537 -112.8551 530.3531 ## 21105 2017-12-09 02:00:40 32.03539 -112.8551 537.5255 ## 22103 2017-12-09 13:00:39 32.03537 -112.8551 531.3355 ``` --- class: inverse, center, middle # R Package Structure --- ## R package structure .pull-left[ 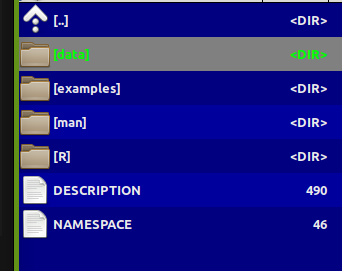 ] .pull-right.large[ - `R/` folder contains code - `data/` folder contains data — as `.rda` - `man/` folder contains documentation - `DESCRIPTION` — essential info - `NAMESPACE` — complicated (*mainly automated*) ] --- ## `DESCRIPTION` file ``` Package: pronghorn Type: Package Title: Sonoran pronghorn analysis project Version: 0.1.0 Author: Elie, Nicki, others Maintainer: The package maintainer <yourself@somewhere.net> Description: The pronghorn package is a PRIVATE collaborative package containing processed data, code and results for analysis of Sonoran pronghorn. License: PRIVATE Encoding: UTF-8 LazyData: false Depends: lubridate, magrittr, plyr, dplyr, ggplot2, ggpubr, sp, sf, stringr Suggests: mapview RoxygenNote: 7.1.1 ``` --- ## Documentation .pull-left[ <img src='images/pronghorn_data1.png' width='100%'> ] .pull-right[ R man file (auto-generated) ``` \docType{data} \name{pronghorn_gps} \title{GPS data of Sonoran pronghorn} \format{ Contains only five columns: \describe{ \item{File}{Original file name} \item{ID}{ID of animal} \item{DateTime}{Date and time in POSIXct} \item{Longitude,Latitude}{} } } \description{ 43 GPS collared pronghorn collared between 2008 and 2020 } ``` ] --- ## Using Roxygen Streamlines documentation by turning "comments" into help files. .small[ ``` r #' GPS data of Sonoran pronghorn #' #' 43 GPS collared pronghorn collared between 2008 and 2020 #' #' @usage data(pronghorn_gps) #' #' @format Contains only five columns: #' \describe{ #' \item{File}{Original file name} #' \item{ID}{ID of animal} #' \item{DateTime}{Date and time in POSIXct} #' \item{Longitude,Latitude}{} #' } #' @example examples/pronghorn_gps_examples.R #' @source Arizona DFG, via Andy Goodwin. #' @keywords data ``` ] > You must install `roxygen2` package and configure build settings (once) to automatically build help files --- class: inverse, center, middle # Building a Package: Step by Step --- class: large ## How to create a package 1. By hand 2. `base::package.skeleton()` 3. `usethis::create_package()` ← **recommended** 4. Build directly off an existing GitHub project --- ## The example: *Paramecium* competition .pull-left.large[ We'll take scripts from [`fittingfunctions.R`](content/fittingfunctions.R) and datasets [`single.csv`](content/single.csv) and [`mixture.csv`](content/mixture.csv) and build a package called `combinator`. It fits a logistic growth model and explores a competition model from Gause's famous 1930s experiment. ] .pull-right[ <img src="Rpackages-slides_files/figure-html/unnamed-chunk-8-1.png" alt="" width="100%" style="display: block; margin: auto;" /> ] --- ## Step I: Build a skeleton (empty) package .pull-left[ Know your working directory first! ``` r getwd() ``` ``` footnotesize ## [1] "C:/Users/egurarie/teaching/EFB654_Materials/2026/20-building-R-packages" ``` Windows users — install Rtools first: ``` r installR::install.Rtools() ``` Create the package skeleton: ``` r require(usethis) create_package("combinator") ``` ] .pull-right[ Output: ``` Package: combinator Title: What the Package Does Version: 0.0.0.9000 Authors@R (parsed): * First Last <first.last@example.com> [aut, cre] Description: What the package does. License: use_mit_license() or friends Encoding: UTF-8 LazyData: true RoxygenNote: 7.1.0 ``` Creates a `combinator/` folder with correct structure and opens a new RStudio project. ] --- ## Step II: Edit the DESCRIPTION file Open and fill in name, license, contact, etc. ``` r use_gpl3_license("combinator") ``` ``` ✓ Setting active project to '.../combinator' ✓ Setting License field in DESCRIPTION to 'GPL-3' ✓ Writing 'LICENSE.md' ✓ Adding '^LICENSE\.md$' to '.Rbuildignore' ``` For a personal-use package, none of this is critical — but good practice. --- ## Step III: Save some data ``` r single <- read.csv("content/single.csv") mixture <- read.csv("content/mixture.csv") ``` Create a `data/` directory and save with `.rda` extension: ``` r save(single, file = "data/single.rda") save(mixture, file = "data/mixture.rda") ``` These can always be loaded independently: ``` r load("data/single.rda") load("data/mixture.rda") ``` --- ## Step IV: Save some code .pull-left-40[ Take these three functions and save each into a **separate** file in `R/`: - `logistic.R` - `fitLogistic.R` - `linesLogistic.R` ] .pull-right-60[ ``` r logistic <- function(x, N0, K, r0){ K / (1 + ((K - N0) / N0) * exp(-r0 * x)) } fitLogistic <- function(data, y = "N", time = "Day", N0 = 1, K = 200, r0 = 0.75){ Y <- with(data, get(y)) X <- with(data, get(time)) myfit <- nls(Y ~ logistic(X, N0, K, r0), start = list(N0 = N0, K = K, r0 = r0)) summary(myfit) } linesLogistic <- function(au.fit, ...){ curve(logistic(x, N0 = au.fit$coefficients[1,1], K = au.fit$coefficients[2,1], r0 = au.fit$coefficients[3,1]), add = TRUE, ...) } ``` ] --- ## Step V: Set up Roxygen ``` r install.packages("roxygen2") ``` Go to `Build > Configure Build Tools > Configure` and check the box next to `Build and Restart`. Modify `logistic.R`: ``` r #' Logistic function #' #' Grows to a carrying capacity K #' #' @param x time #' @param N0 initial population size #' @param K carrying capacity #' @param r0 intrinsic growth rate #' @examples curve(logistic(x, .01, 1, 10)) #' #' @export logistic <- function(x, N0, K, r0){ K / (1 + ((K - N0) / N0) * exp(-r0 * x)) } ``` Every comment leads with `#'`. `@export` makes the function user-accessible. --- ## Step VI: Build the package! Press `Ctrl+Shift+B` or `Build > Clean and Rebuild` ``` Restarting R session... > library(combinator) ``` Type `?logistic` — your first help file! Click `index` at the bottom to see all. .box-green[ **Exercise:** Add title, description, and `@param` tags to `fitLogistic.R` and `linesLogistic.R`, then rebuild. ] --- ## Step VII: Document the data Create `R/datadocumentation.R`: ``` r #' Single separate paramecium growth #' #' Growth of two species of paramecium, P. aurelia and P. caudatum #' #' @usage data(single) #' #' @format Three columns: #' \describe{ #' \item{Day}{Day of experiment} #' \item{caudatum}{volume of P. caudatum} #' \item{aurelia}{volume of P. aurelia} #' } #' #' @examples #' data(single) #' plot(aurelia ~ Day, data = single) #' #' @source Gause (1934) The Struggle for Existence #' @keywords data "single" ``` Rebuild, then try `?single`. --- ## Step VIII: Separate example files Save as `examples/logisticFitExample.R`: ``` r require(combinator) data(single) plot(aurelia ~ Day, data = single, col = 1) points(caudatum ~ Day, data = single, col = 2) fit1 <- fitLogistic(single, y = "aurelia", time = "Day", 1, 200, .75) fit2 <- fitLogistic(single, y = "caudatum", time = "Day", 1, 200, .75) linesLogistic(fit1, lwd = 3) linesLogistic(fit2, col = 2, lwd = 3) ``` Add to `fitLogistic.R`: ``` r #' @example examples/fitLogisticExample.R ``` Note: `@example` (singular) links a script file; `@examples` (plural) takes inline code. --- class: inverse, center, middle # The rest is gravy. <img src='images/gravy.jpeg' width='35%'/> --- class: inverse ## Further resources .large[ - http://web.mit.edu/insong/www/pdf/rpackage_instructions.pdf - https://learn.gencore.bio.nyu.edu/building-r-packages/ - https://bookdown.org/rdpeng/RProgDA/building-r-packages.html ]